There have been a large number of minor enhancements to smooth out workflows and improve compatibility with the macOS and other applications. 
scf files to a project, optionally base call with phred, if needed, then Trim by Quality prior to aligning or assembling. The reads (typically Sanger sequencing reads in ABI or SCF format) should not be aligned prior to running the algorithm. When the Editor or Picture tabs are active, a new floating Graphics Palette is displayed (similar in concept to the Graphics Palette for the single sequence Map tab), allowing you to turn features on and off permitting greater control over how shared domains are displayed and outlined.īoth the Align to Reference and Assembly Project windows now let you trim reads based on quality. There is a new Features tab in the protein multiple sequence alignment window where you can view, edit, create and/or delete features in each of the aligned sequences. 
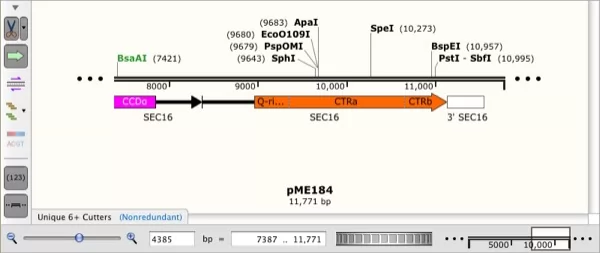
xdna files from Serial Cloner, which is an extended version of the old DNAStrider format.Įnhancements to Outlining Shared Domains in Aligned Sequencesįirst added in MacVector 17.5, this functionality has been significantly enhanced to expose additional Feature-related functionality in Multiple Sequence Alignment documents. 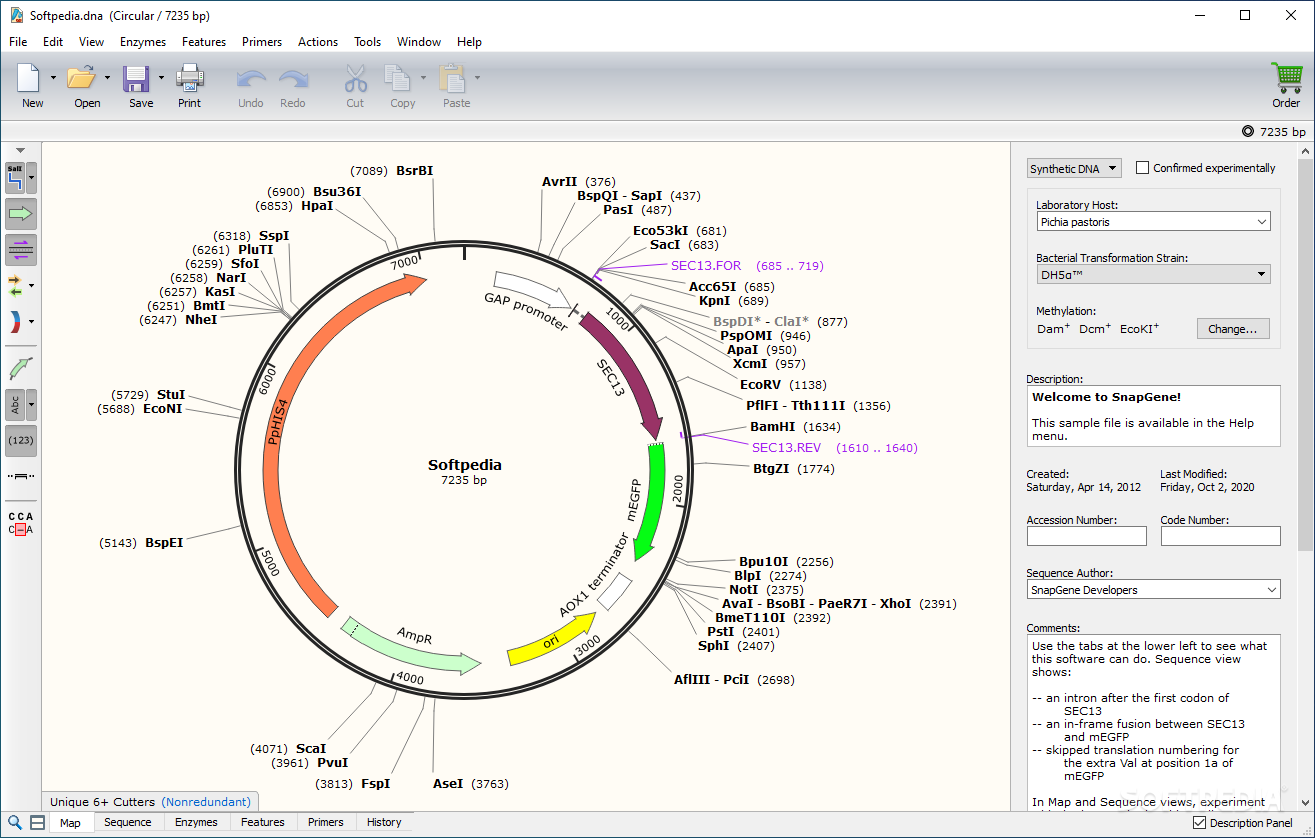
spf Assembly Project files from Sequencher dna files, complete with feature colors, like this plasmid downloaded from the AddGene website The results are shown in the Map tab, and you can zoom in to see the actual sequence of the hits with the guide sequence shown in lower case and the PAM sequence in upper case You can also have MacVector automatically scan for CRISPR PAM sites whenever you open a DNA sequence file - control this using the MacVector | Preferences | Scan DNA pane. The output is a mix between the current Restriction Enzyme and Nucleic Acid Subsequence searches, so you can easily see and copy the guide sequences. While MacVector does not currently search genomes for potential off-target sites, this is still a very useful function to alert you to potential CRISPR target sites. It contains most of the current Dec 2020) characterized Cas9-like enzymes, but you can always open the file and add your own. By default, MacVector uses a file called Protospacer Adjacent Motifs.pam that is located in the / Applications/MacVector/Subsequences/ folder. menu item that will scan Nucleic Acid sequences for Protospacer Adjacent Motifs associated with the CRISPR Cas9 and related enzymes cleavage and modification functions. There is a new feature accessed by the Analyze | CRISPR PAM sites. Otherwise, MacVector 18 has the identical functionality to MacVector 18.1 but with a slightly smaller disk footprint. We recommend this for all users of Intel-base Macintosh computers and for users still unning OS X 10.11 (El Capitan). It requires a minimum of macOS 10.12 (macOS Sierra). MacVector 18.1 is a Universal Binary, meaning that it runs natively on both Intal and Apple Silicon ("M1") Macintosh computers. Finally, there are major enhancements to the protein multiple sequence alignment visualization of domains and the usual macOS compatibility enhancements. There is a new CRISPR PAM sequence searching function and a Trim by Quality option for Sanger sequence files. MacVector 18 adds a number of new functions, including the ability to read a several requested file formats such as Sequencher assembly project files along with Serial Cloner and SnapGene sequence files. Sequence Analysis Tools for Molecular Biologists 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
